An interactive version of this notebook is available on Google Colab or Binder.
Adding a new method#
This tutorial shows how to use the ``add_method`` intervention to introduce a new contraceptive method into an FPsim run at a specified year. You can use it in several ways:
Brand new method: Define a full
fp.Methodand add it, copying switching behavior from an existing method.Quick clone: Clone an existing method (e.g. implants) with minimal config; useful for generic “copy” scenarios.
Clone + override: Clone a method and override specific attributes (name, efficacy, duration, etc.) via
method_pars.Market splitting: Introduce a new method that takes a fraction of another method’s market share using
split_shares.
You always specify when the method becomes available (year) and which existing method to copy switching behavior from (copy_from). Optionally you can pass a custom method, method_pars, and/or split_shares.
Setup#
Import FPsim, Starsim, and set common simulation parameters. We’ll use a small population and short run for quick demos.
[1]:
import fpsim as fp
import starsim as ss
import numpy as np
import matplotlib.pyplot as plt
pars = dict(
n_agents=2000,
start=2000,
stop=2020,
location='kenya',
verbose=0,
analyzers=fp.method_mix_over_time(),
)
intro_year = 2010 # Year when new methods will be introduced
/home/docs/checkouts/readthedocs.org/user_builds/institute-for-disease-modeling-fpsim/envs/latest/lib/python3.11/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
2. Quick clone (copy existing method)#
Add a copy of an existing method without defining a full Method: omit method and use copy_from. The new method is named {source}_copy unless you override it in method_pars.
[3]:
intv_clone = fp.add_method(
year=intro_year,
copy_from='impl',
verbose=True,
)
sim_clone = fp.Sim(pars=pars, interventions=[intv_clone], label='With Implant Clone').run()
Loading data from files in /home/docs/checkouts/readthedocs.org/user_builds/institute-for-disease-modeling-fpsim/envs/latest/lib/python3.11/site-packages/fpsim/locations/kenya/data...
Applying calibration parameters for kenya...
Registered new method "impl_copy" (idx=10), will activate in year 2010
Activating new contraceptive method "impl_copy" in year 2010.0
add_method finalized: "impl_copy" has 119 users (14.90% of method mix)
3. Clone + override with improved contraceptive technology#
Clone an existing method and override specific attributes via method_pars. In this next example we introduce an improved injectable with higher efficacy and a 10% longer relative duration of use.
[4]:
intv_override = fp.add_method(
year=intro_year,
copy_from='inj',
method_pars={
'name': 'inj_improved',
'label': 'Improved Injectable',
'efficacy': 0.998,
'rel_dur_use': 1.1
},
)
sim_override = fp.Sim(pars=pars, interventions=[intv_override], label='With Improved Injectable').run()
Loading data from files in /home/docs/checkouts/readthedocs.org/user_builds/institute-for-disease-modeling-fpsim/envs/latest/lib/python3.11/site-packages/fpsim/locations/kenya/data...
Applying calibration parameters for kenya...
Registered new method "inj_improved" (idx=10), will activate in year 2010
Activating new contraceptive method "inj_improved" in year 2010.0
add_method finalized: "inj_improved" has 87 users (9.80% of method mix)
Alternative: Adding methods from simulation start#
The examples above use the add_method intervention to introduce methods partway through a simulation. If you need a method to be available from the very beginning of the simulation, use the manual method creation approach below.
First, make a copy of the default list of contraceptive methods:
[7]:
import sciris as sc
my_methods = fp.make_methods()
Define a new method with its attributes:
A simple name (used inside the code)
Efficacy (as a decimal)
Whether it is a modern method
Duration of use (in months or as a distribution)
A label (used in plots)
[8]:
new_method = fp.Method(name='new', efficacy=0.96, modern=True, dur_use=15, label='New method')
Add the method to the list:
[9]:
my_methods += new_method
Create a method choice module that includes your new method:
[10]:
method_choice = fp.RandomChoice(methods=my_methods)
Now run simulations comparing baseline (without the new method) to a scenario with the new method:
[11]:
pars = dict(
n_agents = 10_000,
location = 'senegal',
start = 2000,
stop = 2012,
exposure_factor = 1.0 # Overall scale factor on probability of becoming pregnant
)
s1 = fp.Sim(pars=pars, label='Baseline')
s2 = fp.Sim(pars=pars, contraception_module=method_choice, label='New Method')
simlist = sc.autolist([s1, s2])
msim = ss.MultiSim(sims=simlist)
msim.run(parallel=False)
Loading data from files in /home/docs/checkouts/readthedocs.org/user_builds/institute-for-disease-modeling-fpsim/envs/latest/lib/python3.11/site-packages/fpsim/locations/senegal/data...
Applying calibration parameters for senegal...
Loading data from files in /home/docs/checkouts/readthedocs.org/user_builds/institute-for-disease-modeling-fpsim/envs/latest/lib/python3.11/site-packages/fpsim/locations/senegal/data...
Applying calibration parameters for senegal...
Initializing sim "Baseline" with 10000 agents
Running "Baseline": 2000.01.01 ( 0/145) (0.00 s) ———————————————————— 1%
Running "Baseline": 2001.01.01 (12/145) (0.24 s) •——————————————————— 9%
Running "Baseline": 2002.01.01 (24/145) (0.54 s) •••————————————————— 17%
Running "Baseline": 2003.01.01 (36/145) (0.85 s) •••••——————————————— 26%
Running "Baseline": 2004.01.01 (48/145) (1.17 s) ••••••—————————————— 34%
Running "Baseline": 2005.01.01 (60/145) (1.50 s) ••••••••———————————— 42%
Running "Baseline": 2006.01.01 (72/145) (2.01 s) ••••••••••—————————— 50%
Running "Baseline": 2007.01.01 (84/145) (2.34 s) •••••••••••————————— 59%
Running "Baseline": 2008.01.01 (96/145) (2.68 s) •••••••••••••——————— 67%
Running "Baseline": 2009.01.01 (108/145) (3.03 s) •••••••••••••••————— 75%
Running "Baseline": 2010.01.01 (120/145) (3.37 s) ••••••••••••••••———— 83%
Running "Baseline": 2011.01.01 (132/145) (3.73 s) ••••••••••••••••••—— 92%
Running "Baseline": 2012.01.01 (144/145) (4.09 s) •••••••••••••••••••• 100%
Initializing sim "New Method" with 10000 agents
Running "New Method": 2000.01.01 ( 0/145) (0.00 s) ———————————————————— 1%
Running "New Method": 2001.01.01 (12/145) (0.14 s) •——————————————————— 9%
Running "New Method": 2002.01.01 (24/145) (0.34 s) •••————————————————— 17%
Running "New Method": 2003.01.01 (36/145) (0.53 s) •••••——————————————— 26%
Running "New Method": 2004.01.01 (48/145) (0.73 s) ••••••—————————————— 34%
Running "New Method": 2005.01.01 (60/145) (0.93 s) ••••••••———————————— 42%
Running "New Method": 2006.01.01 (72/145) (1.13 s) ••••••••••—————————— 50%
Running "New Method": 2007.01.01 (84/145) (1.33 s) •••••••••••————————— 59%
Running "New Method": 2008.01.01 (96/145) (1.54 s) •••••••••••••——————— 67%
Running "New Method": 2009.01.01 (108/145) (1.75 s) •••••••••••••••————— 75%
Running "New Method": 2010.01.01 (120/145) (1.96 s) ••••••••••••••••———— 83%
Running "New Method": 2011.01.01 (132/145) (2.17 s) ••••••••••••••••••—— 92%
Running "New Method": 2012.01.01 (144/145) (2.38 s) •••••••••••••••••••• 100%
[11]:
MultiSim("Baseline"; n_sims: 2; base: Sim(Baseline; n=10000; 2000—2012; demographics=deaths; connectors=contraception, edu, fp))
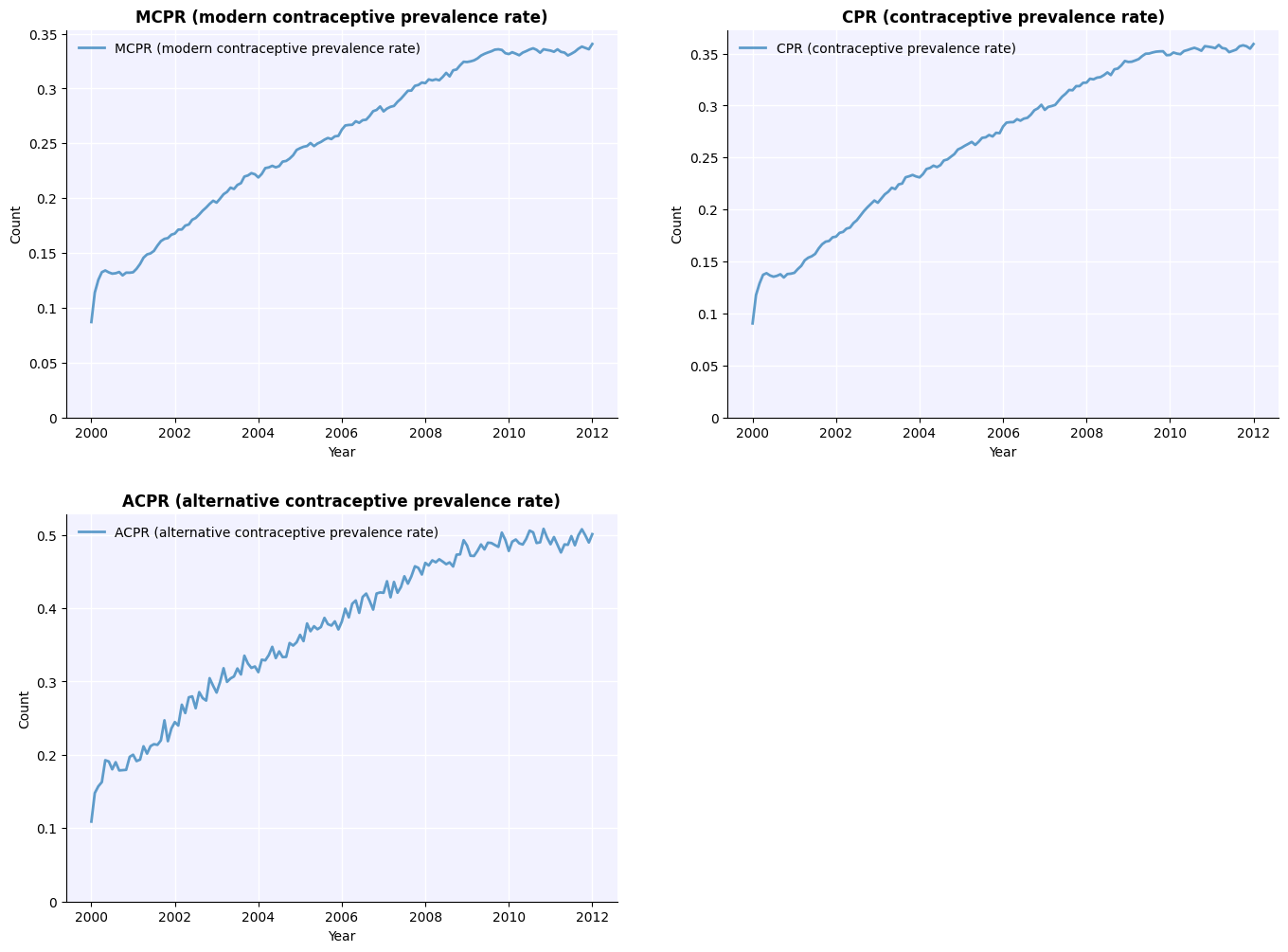
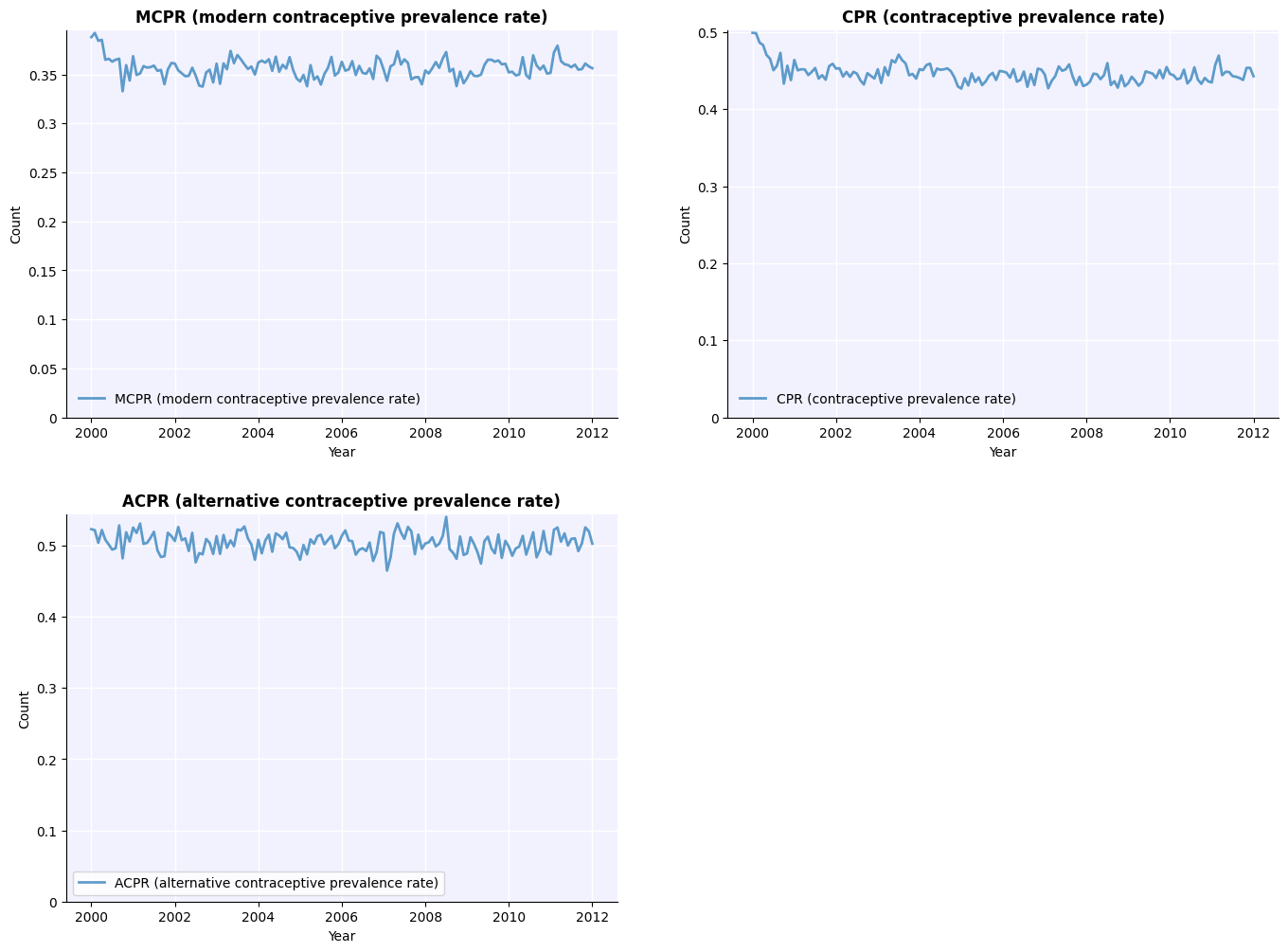
Compare the contraceptive prevalence rate between the two scenarios:
[12]:
msim.plot(key='cpr');
Figure(1600x1000)