T11 - Advanced features#
This tutorial covers advanced features of Covasim, including custom population options and changing the internal computational methods.
Click here to open an interactive version of this notebook.
Defining populations with SynthPops#
For complex populations, we suggest using SynthPops, a Python library designed specifically for this purpose. In contrast the population methods built-in to Covasim, SynthPops uses data to produce synthetic populations that are statistically indistinguishable from real ones. For a relatively complex example of how SynthPops was used to create a complex school network for the Seattle region, see here.
Defining contact layers#
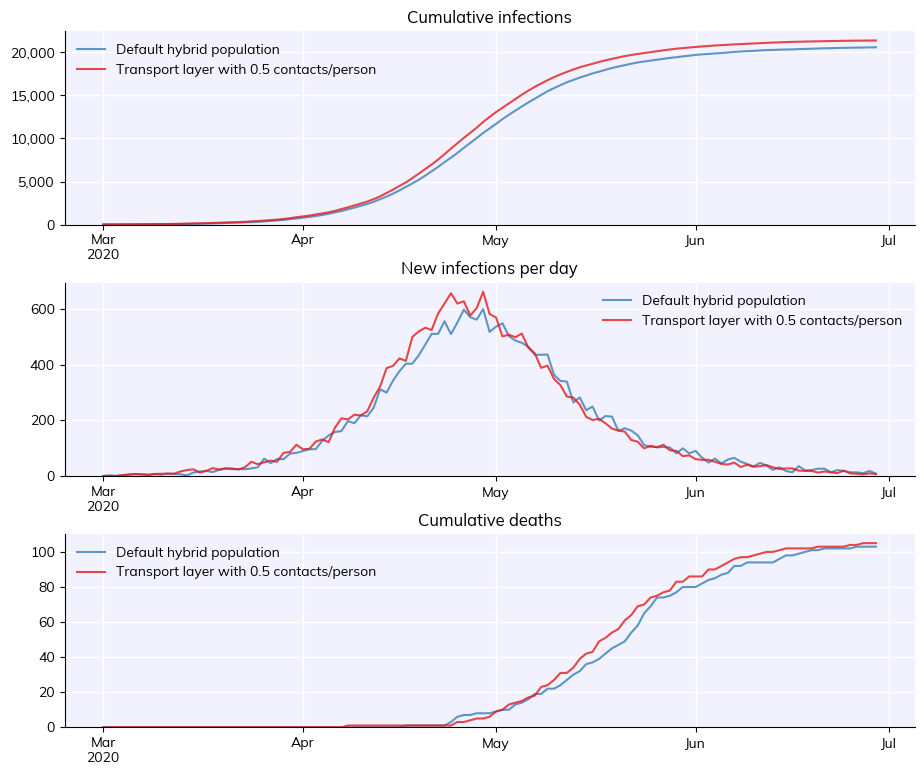
As mentioned in Tutorial 4, contact layers are the graph connecting the people in the simulation. Each person is a node, and each contact is an edge. While enormous complexity can be used to define realistic contact networks, a reasonable approximation in many cases is random connectivity, often with some age assortativity. Here is an example for generating a new contact layer, nominally representing public transportation, and adding it to a simulation:
[1]:
import numpy as np
import covasim as cv
cv.options(jupyter=True, verbose=0)
# Create the first sim
orig_sim = cv.Sim(pop_type='hybrid', n_days=120, label='Default hybrid population')
orig_sim.initialize() # Initialize the population
# Create the second sim
sim = orig_sim.copy()
# Define the new layer, 'transport'
n_people = len(sim.people)
n_contacts_per_person = 0.5
n_contacts = int(n_contacts_per_person*n_people)
contacts_p1 = cv.choose(max_n=n_people, n=n_contacts)
contacts_p2 = cv.choose(max_n=n_people, n=n_contacts)
beta = np.ones(n_contacts)
layer = cv.Layer(p1=contacts_p1, p2=contacts_p2, beta=beta) # Create the new layer
# Add this layer in and re-initialize the sim
sim.people.contacts.add_layer(transport=layer)
sim.reset_layer_pars() # Automatically add layer 'transport' to the parameters using default values
sim.initialize() # Reinitialize
sim.label = f'Transport layer with {n_contacts_per_person} contacts/person'
# Run and compare
msim = cv.parallel(orig_sim, sim)
msim.plot()
Covasim 3.1.7 (2026-02-18) — © 2020-2026 by IDM
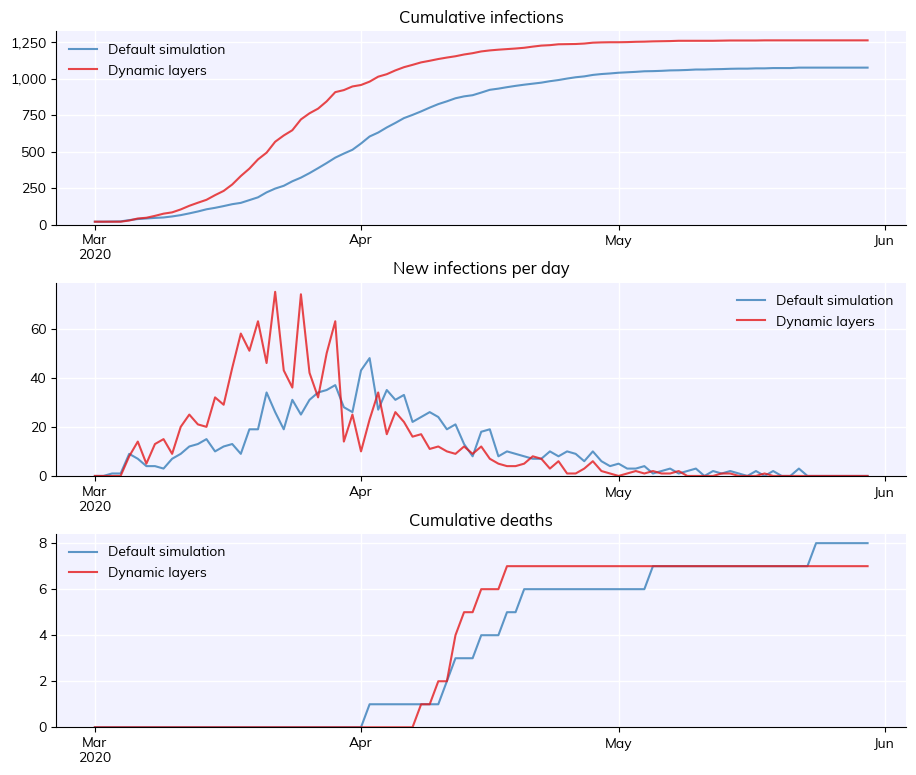
Defining dynamic layers#
You can also define custom layers that update dynamically, e.g. based on a supplied number of contacts per day. To do this, create a new Layer class and define the update() method. For example:
[2]:
import covasim as cv
import numpy as np
import sciris as sc
class CustomLayer(cv.Layer):
''' Create a custom layer that updates daily based on supplied contacts '''
def __init__(self, layer, contact_data):
''' Convert an existing layer to a custom layer and store contact data '''
for k,v in layer.items():
self[k] = v
self.contact_data = contact_data
return
def update(self, people):
''' Update the contacts '''
pop_size = len(people)
n_new = self.contact_data[people.t] # Pull out today's contacts
self['p1'] = np.array(cv.choose_r(max_n=pop_size, n=n_new), dtype=cv.default_int) # Choose with replacement
self['p2'] = np.array(cv.choose_r(max_n=pop_size, n=n_new), dtype=cv.default_int) # Paired contact
self['beta'] = np.ones(n_new, dtype=cv.default_float) # Per-contact transmission (just 1.0)
return
# Define some simple parameters
pars = sc.objdict(
pop_size = 1000,
n_days = 90,
)
# Set up and run the simulation
base_sim = cv.Sim(pars, label='Default simulation')
sim = cv.Sim(pars, dynam_layer=True, label='Dynamic layers')
sim.initialize()
# Update to custom layer
for key in sim.layer_keys():
contact_data = np.random.randint(pars.pop_size*10, pars.pop_size*20, size=pars.n_days+1) # Generate a number of contacts for today
sim.people.contacts[key] = CustomLayer(sim.people.contacts[key], contact_data=contact_data)
# Run and plot
msim = cv.parallel(base_sim, sim)
msim.plot()
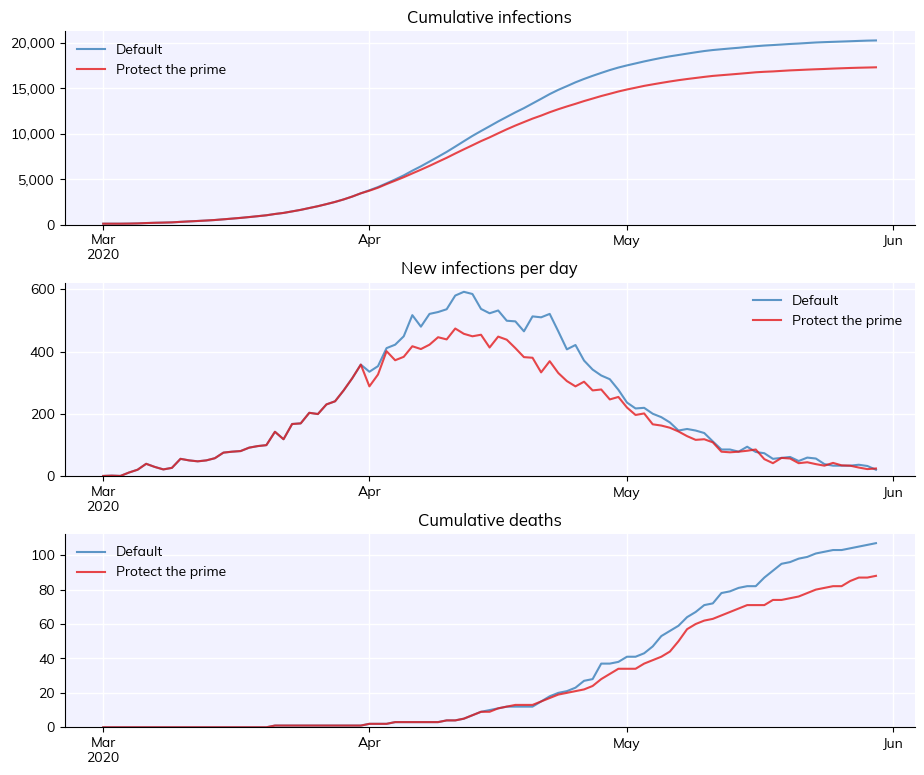
Defining custom population properties#
Another useful feature is adding additional features to people, for use in subtargeting. For example, this example shows how to define a subpopulation with higher baseline mortality rates. This is a simple example illustrating how you would identify and target people based on whether or not the have a prime-number index, based on the protecting the elderly example from Tutorial 1.
[3]:
import numpy as np
import sciris as sc
import covasim as cv
def protect_the_prime(sim):
if sim.t == sim.day('2020-04-01'):
are_prime = sim.people.prime
sim.people.rel_sus[are_prime] = 0.0
pars = dict(
pop_type = 'hybrid',
pop_infected = 100,
n_days = 90,
verbose = 0,
)
# Default simulation
orig_sim = cv.Sim(pars, label='Default')
# Create the simulation
sim = cv.Sim(pars, label='Protect the prime', interventions=protect_the_prime)
sim.initialize() # Initialize to create the people array
sim.people.prime = np.array([sc.isprime(i) for i in range(len(sim.people))]) # Define whom to target
# Run and plot
msim = cv.parallel(orig_sim, sim)
msim.plot()
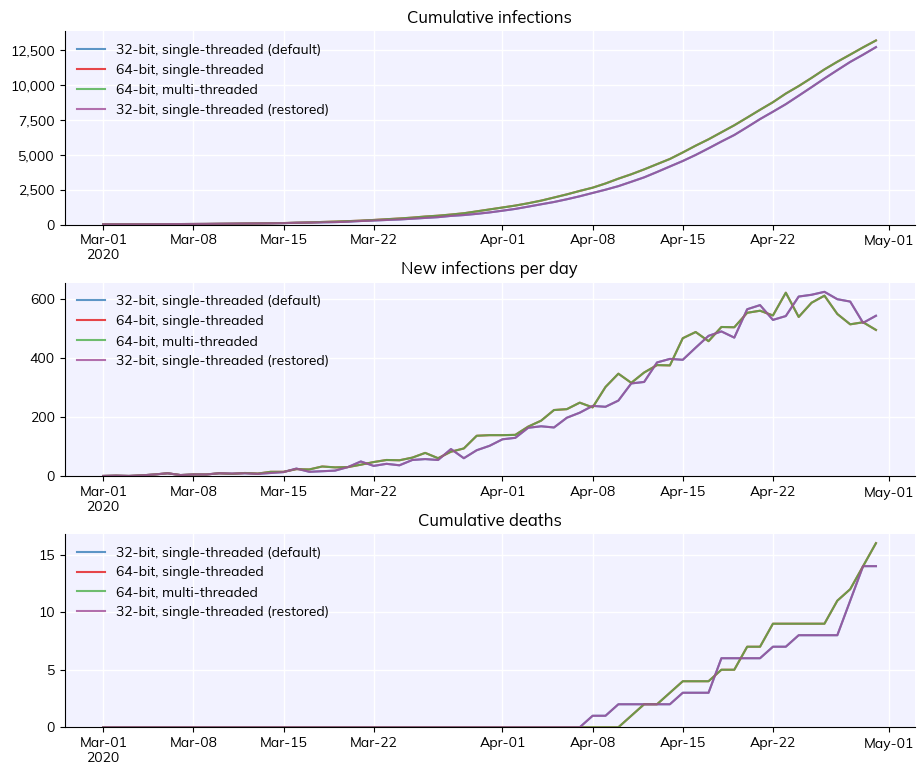
Changing Numba options#
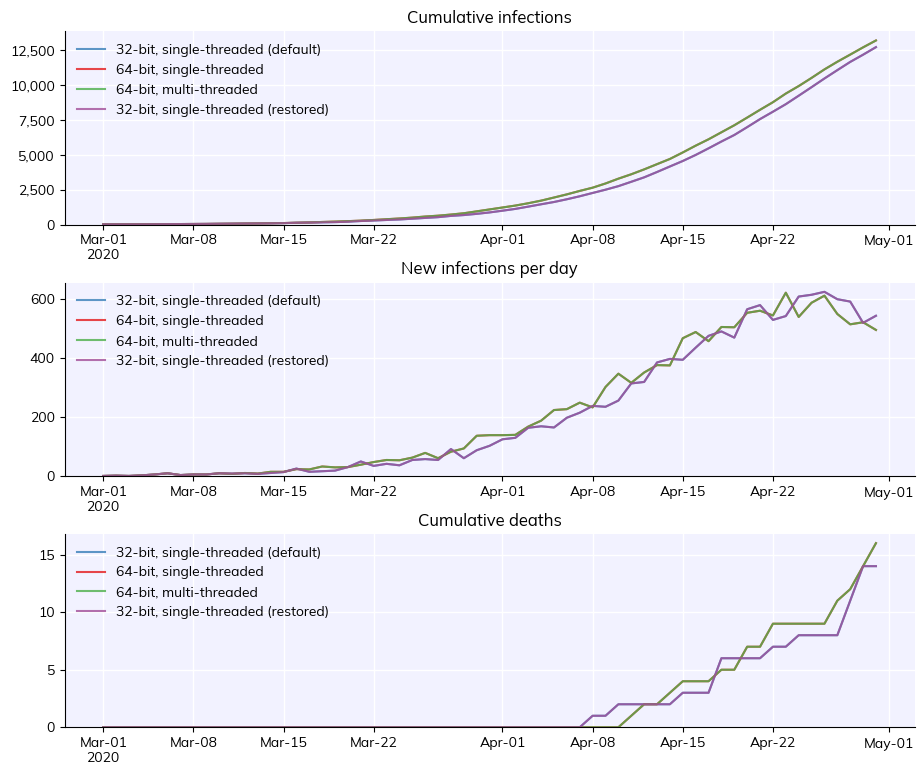
Finally, this example shows how you can change the default Numba calculation options. It’s not recommended – especially running with multithreading, which is faster but gives stochastically unreproducible results – but it’s there if you want it.
[4]:
import covasim as cv
# Create a standard 32-bit simulation
sim32 = cv.Sim(label='32-bit, single-threaded (default)', verbose='brief')
sim32.run()
# Use 64-bit instead of 32
cv.options.set(precision=64)
sim64 = cv.Sim(label='64-bit, single-threaded', verbose='brief')
sim64.run()
# Use parallel threading
cv.options.set(numba_parallel=True)
sim_par = cv.Sim(label='64-bit, multi-threaded', verbose='brief')
sim_par.run()
# Reset to defaults
cv.options.set('defaults')
sim32b = cv.Sim(label='32-bit, single-threaded (restored)', verbose='brief')
sim32b.run()
# Plot
msim = cv.MultiSim([sim32, sim64, sim_par, sim32b])
msim.plot()
Sim("32-bit, single-threaded (default)"; 2020-03-01 to 2020-04-30; pop: 20000 random; epi: 12730⚙, 14☠)
Reloading Covasim so changes take effect...
Reload complete. Note: for some options to take effect, you may also need to delete Covasim's __pycache__ folder.
Sim("64-bit, single-threaded"; 2020-03-01 to 2020-04-30; pop: 20000 random; epi: 13209⚙, 16☠)
Reloading Covasim so changes take effect...
Reload complete. Note: for some options to take effect, you may also need to delete Covasim's __pycache__ folder.
Sim("64-bit, multi-threaded"; 2020-03-01 to 2020-04-30; pop: 20000 random; epi: 13209⚙, 16☠)
Reloading Covasim so changes take effect...
Covasim 3.1.7 (2026-02-18) — © 2020-2026 by IDM
Reload complete. Note: for some options to take effect, you may also need to delete Covasim's __pycache__ folder.
Sim("32-bit, single-threaded (restored)"; 2020-03-01 to 2020-04-30; pop: 20000 random; epi: 12730⚙, 14☠)
[4]: